R.1: Another In The Growing List Of SARS-CoV-2 Variants
(Posted on Monday, September 20, 2021)

Welcome to Kentucky road sign at the state border
GETTY
Update: This article was edited to mention that R.1 is no longer spreading significantly in the United States. This version highlights the R.1 variant’s capability to initiate breakthrough infection in fully-vaccinated elderly populations.
A variant detected in a Kentucky nursing home in March 2021, infecting 45 residents and health care personnel. Many fell ill and one died. The variant, which was first detected in Japan in December 2020, has over 10,000 entries in the GISAID SARS-CoV-2 database. In the United States, R.1 infections peaked in May 2021 and were last detected in early August 2021. This variant, like many others, was displaced by the Delta variant, which is now the cause of nearly all infections in the United States. We focus on this variant because unlike most variants of concern, the R.1 variant has relatively few mutations in the Spike protein in proportion to mutations in other regions of the genome. For this reason, it is worth understanding the mutations in virus in more detail.
The variant contains five mutations previously noted in variants of concern or interest, two of which are in the Spike protein (Figure 1). It also contains many unique mutations. Here we describe the potential effects of each mutation on replication, immune evasion, and pathogenesis.
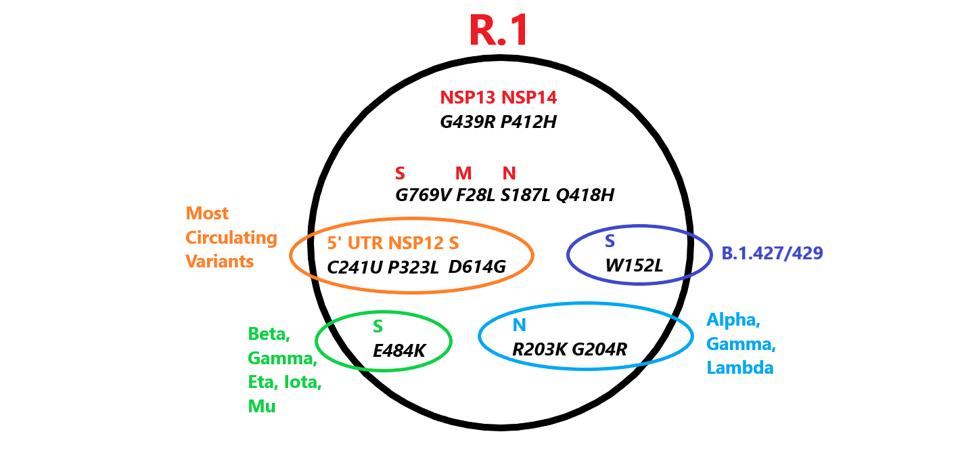
FIGURE 1: Mutations in R.1 found in other variants. Those in red are unique to R.1.
ACCESS HEALTH INTERNATIONAL
The Triad
The R.1 variant shares a common origin with all variants of interest or concern. They are marked by what I call the Triad, three mutations, one the 5’ untranslated region: C241U, a second in the viral polymerase NSP12: P323L, and the third D614G in the exterior S1 domain of the spike protein. The D614G mutation increases the infectivity. The contribution of the other two members of the Triad remains a mystery. Together these three mutations characterize the first major variant first observed in early 2020. That virus soon displaced almost all of the original Wuhan isolates. The Triad virus is the parent of all variants of concern or, interest including Alpha, Beta, Gamma, Delta, Lambda, and Mu. Only two currently circulating variants, A.30 and A.23.1, both of East Africa origin, lack the Triad and are most likely independent descendants of the original Wuhan virus.
The Spike Protein (S)
The spike protein of SARS-CoV-2 binds the host ACE2 receptor and initiates infection. The R.1 variant contains three Spike mutations, two of which are previously noted in other variants the other which is unique (Figure 2).
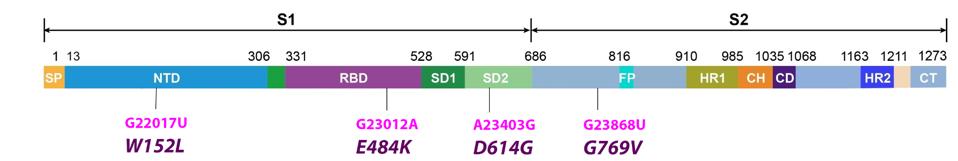
FIGURE 2: Common mutations in the nonstructural proteins in the R.1 variant genome. Color code: nonsynonymous mutations (pink) and their corresponding amino acid mutations (purple).
ACCESS HEALTH INTERNATIONAL
One of the frequently mutated domains of the spike protein is the receptor-binding domain. The R.1 variant contains one mutation in this domain, E484K. The 484 mutation conveys increased resistance to antibodies in convalescent sera and to neutralizing monoclonal antibodies. The E484K substitutes the negatively charged residue glutamic acid for the positively charged lysine. The E484K is present in the Beta, Gamma, Eta, Iota, and Mu variants.
R.1 contains the W152L mutation in the N-terminal domain. This region of the spike protein is also the target of neutralizing antibodies in convalescent sera and is the target of several neutralizing monoclonal antibodies. An identical mutation at position 152 is present in one of the minor variants of the Delta strain found in India. The very same amino acid is mutant in several variants first detected in California, B.1427/429. The 152 tryptophan is substituted for leucine in the California variants. Multiple independent mutations of the same amino acid imply functional advantage.
The mutation, G769V, falls in an interdomain region between positions 732 and 780. It is unique to the R.1 variant.
The Non-Structural Proteins
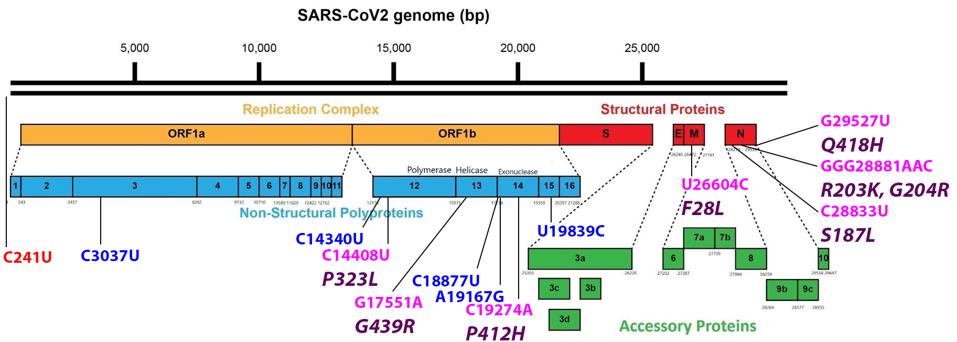
FIGURE 3: Common mutations external to the S protein in the R.1 variant genome. Color code: noncoding nucleotide mutations (red), synonymous mutations (blue), nonsynonymous mutations (pink), and their corresponding amino acid mutations (purple).
ACCESS HEALTH INTERNATIONAL
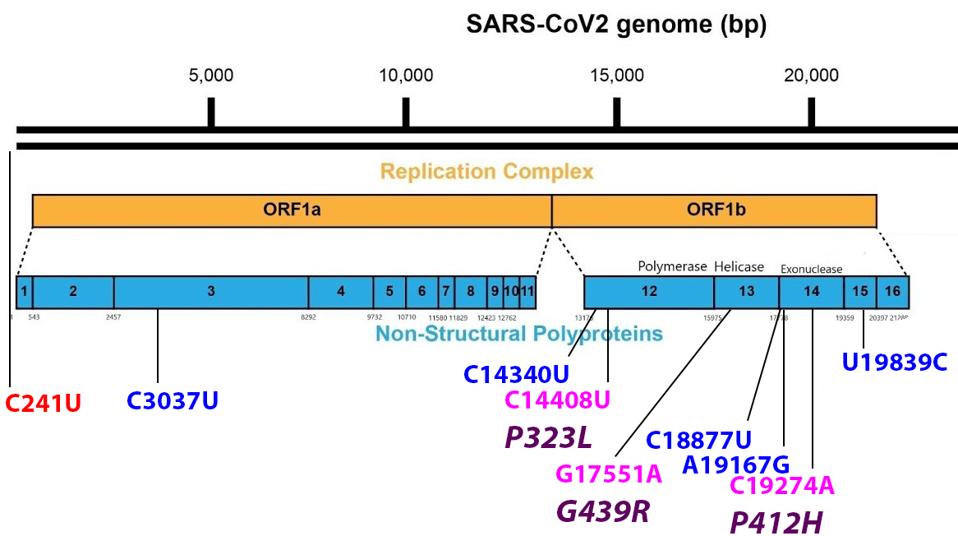
FIGURE 4: Mutations found in the Orf1ab non-structural proteins. Color code: noncoding nucleotide mutations (red), synonymous mutations (blue), nonsynonymous mutations (pink), and their corresponding amino acid mutations (purple).
ACCESS HEALTH INTERNATIONAL
The Non-Structural Protein 13 (NSP13)
The NSP13 protein is the helicase and part of the replication transcription complex. The G439R mutation of R.1 is unique to this variant. The 439 residue lies adjacent to an active ATP-binding site (Figure 5 and 6). The Glycine to arginine mutation is a change from a neutral amino acid to one that carries a positive charge. The effect of this mutation on replication and helical activity is worth further study.
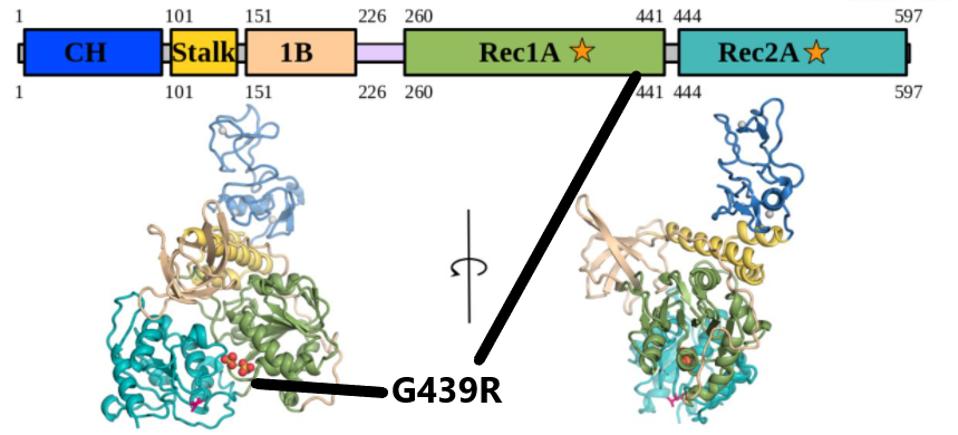
FIGURE 5: The G439R mutation in NSP13. The mutation falls in the Rec1A domain, noted by the black lines.
WHITE ET AL
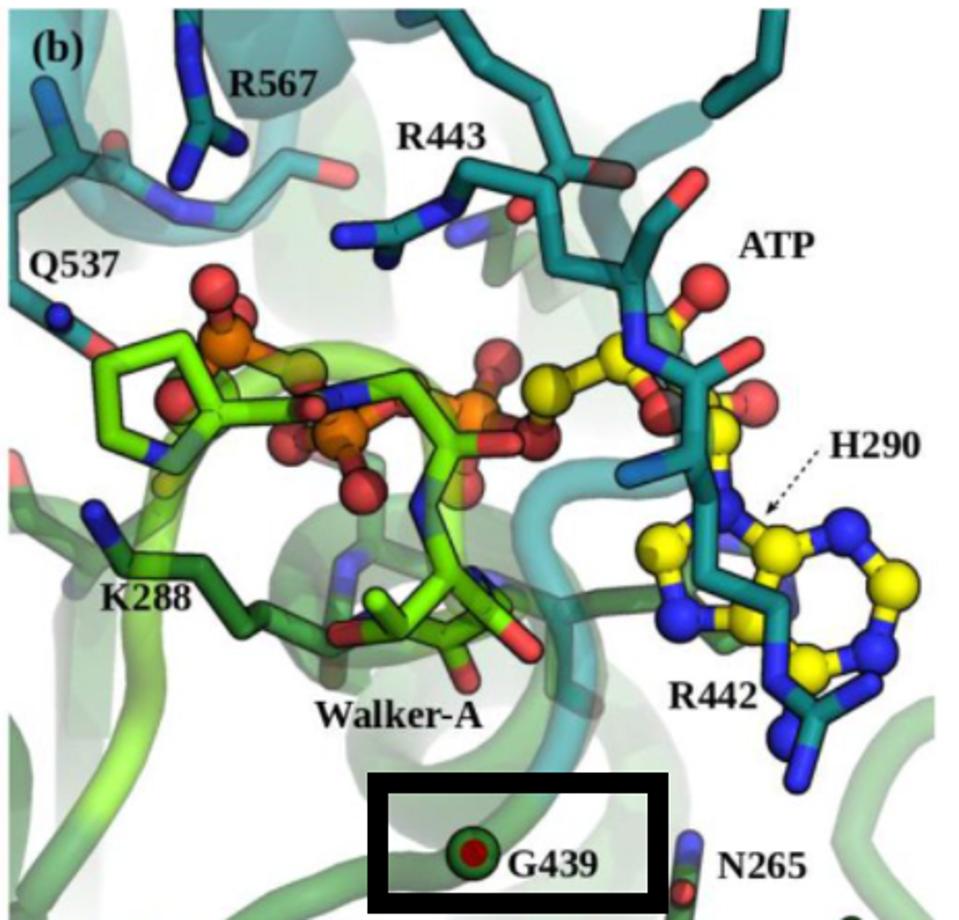
FIGURE 6: Position G439 located near the ATP-binding site in NSP13.
WHITE ET AL
The Non-Structural Protein 14 (NSP14)
NSP14 is an exoribonuclease that, like the NSP13 helicase, is a part of the replication transcription complex. The exoribonuclease has an error-editing function and also confers resistance to chain-terminating nucleotides. The NSP14 protein of R.1 carries the unique P412H mutation. NSP14 is also is a methyltransferase that participates in the formation of capped viral messenger RNAs. The capping of viral messenger RNAs is required to circumvent triggering the innate immune response. A change from the amino acid proline to histidine substitutes a neutral amino acid for one that carries a positive charge, again indicating a major charge shift and may enhance the properties of the protein. The site of the mutation lies within the N7 methyltransferase capping domain of the protein adjacent to SAH and GpppA binding (Figure 7).
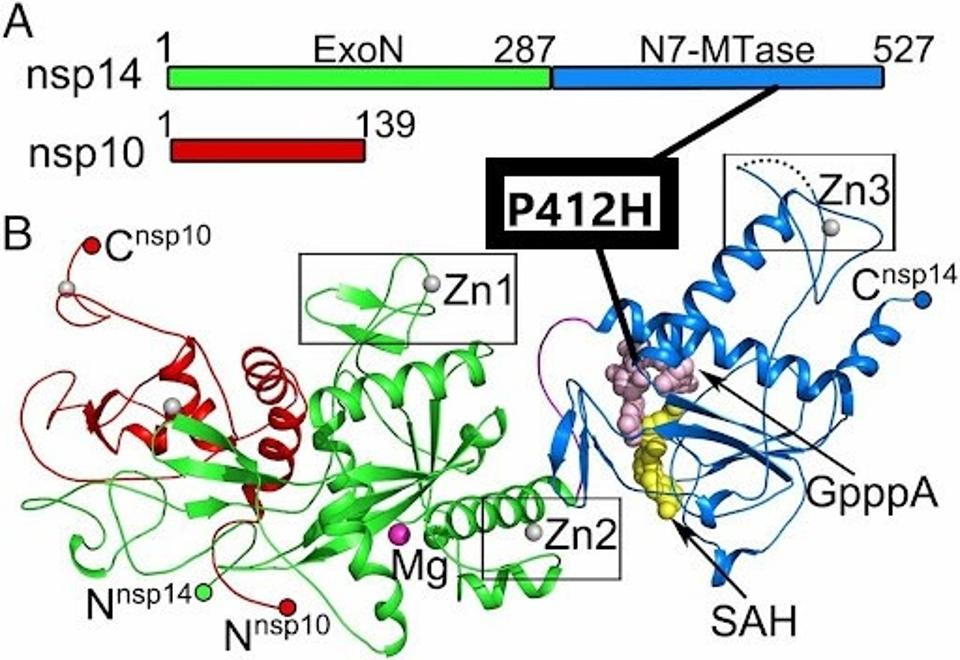
FIGURE 7: The P412H mutation in NSP14. The mutation is located in the N7-MTase subdomain near SAH and GpppA binding sites.
YUANYUAN ET AL
We also observe a number of synonymous nucleotide changes, which could impact the RNA replication and transcription processes. In the nonstructural proteins, we note C3037U, C14340U, C18877U, A19167G, and T19839C. It is possible that, while these do not change amino acids, they could change the secondary structure of the RNA which may result in functional consequences.
The Structural Proteins
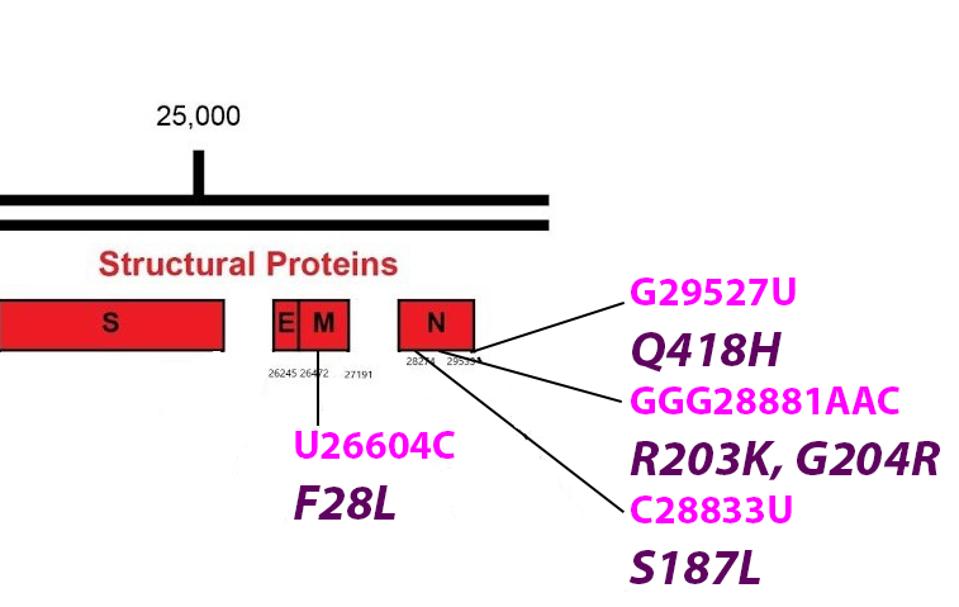
FIGURE 8: Common mutations in the non-Spike structural proteins in the R.1 variant genome. Color code: noncoding nucleotide mutations (red), synonymous mutations (blue), nonsynonymous mutations (pink), and their corresponding amino acid mutations
ACCESS HEALTH INTERNATIONAL
The Membrane Protein (M)
The Membrane protein is key to the formation of virus particles. It is embedded in the membrane that surrounds the virus. M is reported to assist in the packaging of viral into mature virions. M also is active in the suppression of innate immunity. The F28L is unique to R.1. The amino acid change lies within the first of three transmembrane regions domains. Phenylalanine and leucine are both nonpolar and uncharged amino acids. Nonetheless, the variant could impact the multiple functions of M including the stability of the virus particle.
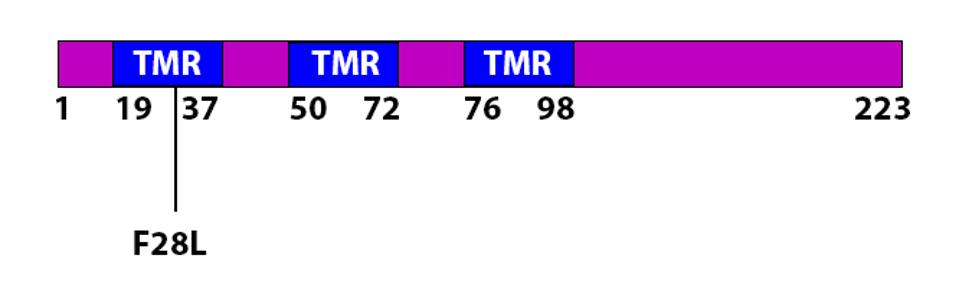
FIGURE 9: The F28L mutation in the M protein. The mutation is found in a transmembrane region (TMR).
ACCESS HEALTH INTERNATIONAL
The Nucleocapsid Protein (N)
The N is the most active of the virus structural proteins. N wraps the viral genome into a helical structure and is required for replication and transcription. During early infection, N is reported to play an important role in the suppression of innate immunity. N is highly antigenic and most people infected people have significant titers of anti-N antibodies.
R.1 is mutant at four positions with N. R.1 shares the R203K and G204R mutations with three variants of concern Alpha, Gamma, and Lambda and with other reported variants as well. The amino acids change in the neighboring amino acid at positions 203 and 204 are always found as a pair. The two mutations lie near the center of the linker domain that connects the RNA binding domain of N with the dimerization domain. The change from glycine to arginine at position 204 introduces a second positive charge into a region that is predominantly neutral.
One of the N mutations S187L is also located in the linker domain. The Q418H adds a positive charge to the extreme carboxy terminus of N.
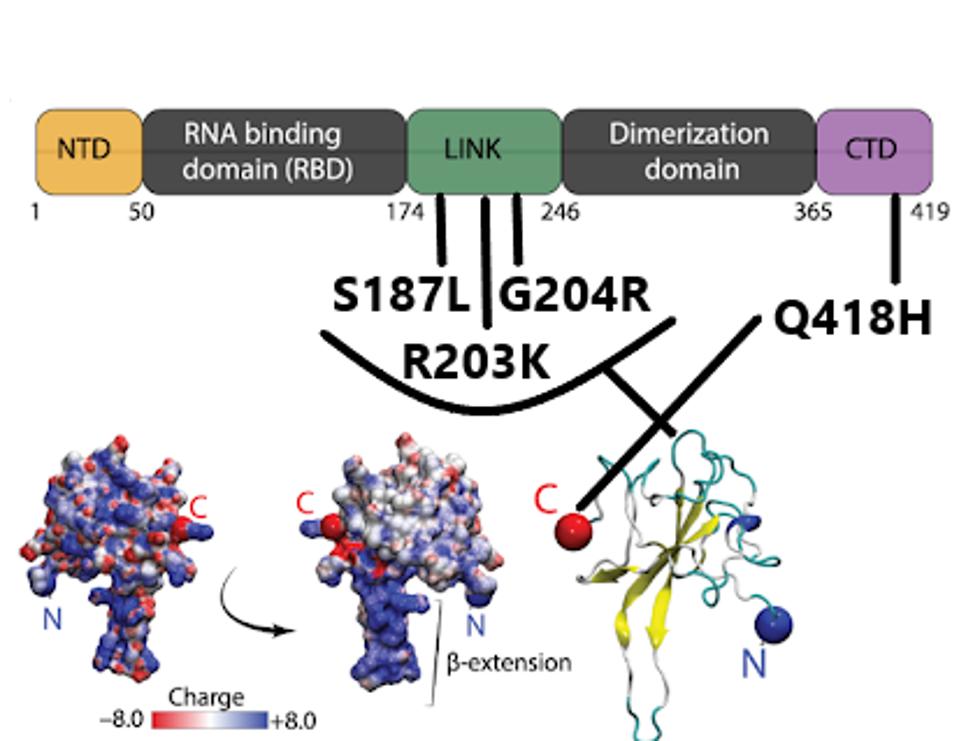
FIGURE 10: R.1 mutations in the N protein, found in the linker and C-terminal domain, followed by a 3D representation.
CUBUK ET AL
R.1 is a variant to watch. As the Delta variant subsides, other mutant strains wait in the wings to potentially drive the next wave of infections in the months to come. In addition to several mutations notably in the spike and nucleocapsid protein in common with variants of concern, R.1 has a set of unique mutations that may confer an additional advantage in transmission, replication, and immune suppression. While R.1 has grown relatively quiet in the GISAID database of late, its mutation-driven properties may propel it to reemerge in the future.
Read the article on Forbes, originally published on September 20th, 2021.

