New Nigerian Variant Continues The Trend Of Dangerous Strains Threatening Covid-19 Progress
(Posted on Wednesday, February 24, 2021)
Random variation is an essential component of all living things. It drives diversity, and it is why there are so many different species. Viruses are no exception. Most viruses are experts at changing genomes to adapt to their environment. We now have evidence that the virus that causes Covid, SARS-CoV-2, not only changes but changes in ways that are significant. This is the twenty-first part of a series of articles on how the virus changes and what that means for humanity. Read the rest: part one, part two, part three, part four, part five, part six, part seven, part eight, part nine, part ten, part eleven, part twelve, part thirteen, part fourteen, part fifteen, part sixteen, part seventeen, part eighteen, part nineteen, and part twenty.
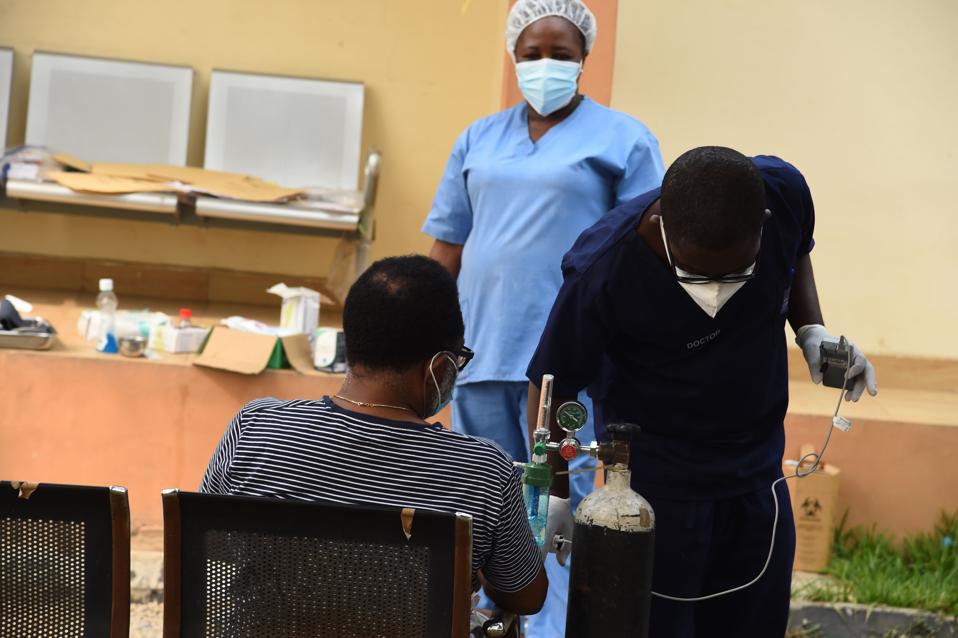
Health officials attend to a COVID-19 patient receiving oxygen therapy at the Infectious Disease Hospital (IDH) in Yaba, Lagos, on January 22, 2021. – The number of confirmed COVID-19 cases in Nigeria is increasing rapidly following daily reports by
AFP VIA GETTY IMAGES
The near-simultaneous detection of SARS-CoV-2 variants from the four corners of the globe deepens our uncertainty regarding the pandemic’s future and our ability to control Covid-19. How serious the problem will be will depend on how many different variants there are, how they differ from one another, how capable they are of reinfecting people, and resisting current and future vaccines.
In this context, here we describe the newfound Nigerian variant (B.1.525). The Nigerian isolates carry mutations mirroring those seen in previous variants and some unique to their structure.
B.1.525 was first detected by genome sequence in mid-December in Nigeria but was also quickly found in cases in the United Kingdom, France, and elsewhere. After only two months, B.1.525 represented over 20% of genomes sequenced in Nigeria. It has been detected in more than 200 cases globally.
The mutation of B.1.525 may increase transmissibility, virulence, and immune escape. The amino acid substitutions Q52R and A67V are located in the spike protein’s N-terminal domain, as are the deletions at positions 69-70 and 144. (See figure). The two point mutations are unique to the B.1.525. Both deletions are characteristics of the UK variant B.1.1.7. The deletion of 69-70 is also found in several naturally occurring variants. No specific function is as yet assigned to the N-terminal domain. Many mutations (and even multiple deletions) in this region are not deleterious to either the transmission, replication, or virulence. The N-terminal domain is highly antigenic. The primary effect of mutations in this region is likely neutralization escape.
The amino acid substitution at position 888 is in the membrane-proximal stem region of the spike protein, nearby a similar change at position 870 found in a neutralization escape variant isolated from a person with persistent SARS-CoV-2 infection.
Among the point mutations and deletions in the spike protein, B.1.525 carries several ubiquitous in SARS-CoV-2 variants today. D614G confers increased transmissibility and is present in nearly all SARS-CoV-2 variants at this stage. The mutation E484K is also present in the South African and Brazilian strains. Amino acid 484 is located in the receptor-binding motif. The E484K change confers approximately three-fold resistance to several Covid vaccines and tenfold resistance to most convalescent antibodies.
The amino acid substitution at position 677 ( Q677H) is identical to that found in variants recently described in the United States. These mutations occur in the S1 portion of the spike protein found far from the receptor-binding domain. Like the D614G mutation (also present in the Nigerian variant), the change at position 677 may increase the stability of the S1-S2 association and thereby increase transmissibility. The 677 mutations may also favor the open rather than closed conjugation of the receptor-binding domain.
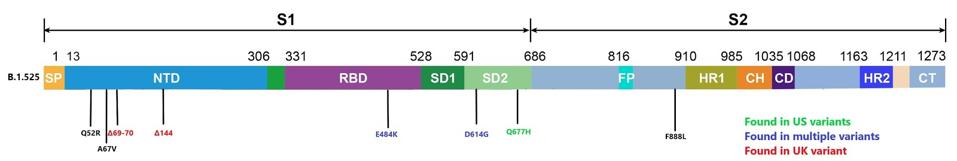
Mutations along the spike protein of the Nigerian variant.
The Nigerian variant NB.1.525 is also rich in mutations in the genome regions that specify proteins other than the spike (Figure 2.) Three of these occur in the genes that comprise the massive replication complex, including three in Orf1a NSP3: (T1189I), NSP6: (a three amino acid deletion 106- 108), and one in Orf1b NSP12:(P323F)- the major RNA dependent RNA polymerase.
Four other mutations occur in the genes specifying viral structural proteins E: (l21F), M:(I82 F), and N:(A12G) and (T208I). It is noteworthy most of these mutations occur in transmembrane proteins NSP6, E, M, and that the N protein is highly antigenic in people infected by SARS-CoV-2. All these mutations may assist neutralization escape. The remaining variants in orf1a may increase replication efficiency, immune escape, or both.
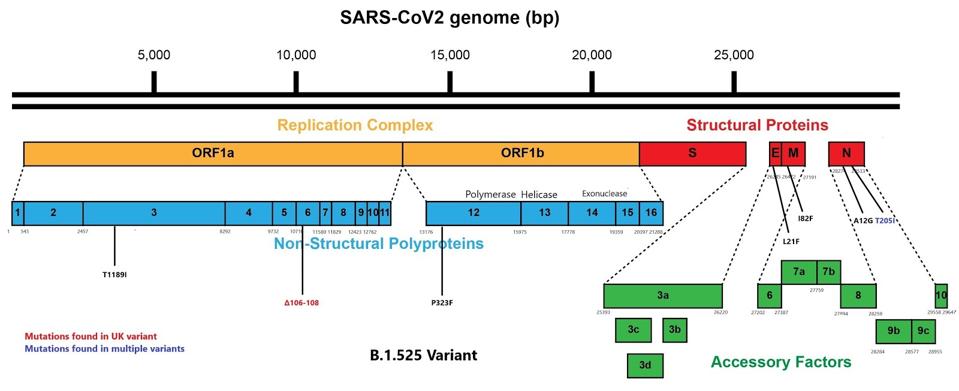
Graphic depicting mutations to Nigerian variant.
This variant maintains the trend we have seen in the past few months. As genome sequencing efforts increased, more new variants are discovered. With each new variant comes more nuanced mutations to its structure that aid its proliferation. Some of the mutations and variants spreading at this very moment confer vaccine-resistance.
I fear a relaxation of public health measures in reaction to vaccine distribution and recent declination in Covid-19 cases. Understandably, the ordinary person thinks themself safe after receiving their vaccine. Antibodies, however, do not last forever, and if new variants are immune-resistant, they may not even be fully effective.
With each new variant, we gain increased insight into the multiple ways SARS-CoV-2 is adapting to us, its host. These include:
Increasing the avidity of the virus for the receptor;
Increasing transmissibility;
Evasion of both novel and existing immune defenses;
Increase replication competence;
Increase in the duration of productive viremia.
B.1.525 now joins the growing array of variants of concern. The protean ability of SARS-CoV-2 to circumvent social and medical containment strategies raises serious questions we must now adapt to this new reality.
Read the full article on Forbes.
Originally published on February 24, 2021.

