This Region Of The Covid-19 Virus Is One We Can’t Ignore
(Posted on Wednesday, February 17, 2021)
Random variation is an essential component of all living things. It drives diversity, and it is why there are so many different species. Viruses are no exception. Most viruses are experts at changing genomes to adapt to their environment. We now have evidence that the virus that causes Covid, SARS-CoV-2, not only changes, but changes in ways that are significant. This is the thirteenth part of a series of articles on how the virus changes and what that means for humanity. Read the rest: part one, part two, part three, part four, part five, part six, part seven, part eight, part nine, part ten, part eleven, and part twelve.

Colorized scanning electron micrograph of a VERO E6 cell (blue) heavily infected with SARS-COV-2 virus particles (orange), isolated from a patient sample. Image captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort
BSIP/UNIVERSAL IMAGES GROUP VIA GETTY IMAGES
In my last two articles, I went over the case studies of the London and Boston patients. Both were immunocompromised and persistently infected with Covid-19. Both also had, in the samples of SARS-CoV-2 taken from their bodies by researchers, mutations that arose independently, but were identical to those seen in new variants of the virus—a phenomenon previously documented by Jesse Bloom and his colleagues in influenza viruses. These cases and many similar to theirs, I argued, open up a window into what this virus has in store for us.
Which brings me to my next subject, one I’ll call the Pittsburgh patient, whose story was documented in a study published in Science magazine earlier this month. Prior to their positive Covid-19 diagnosis, this patient was being treated for cancer and, the authors of the study note, already immunocompromised. The Pittsburgh patient endured severe Covid-19 for about two and a half months before succumbing to the disease.
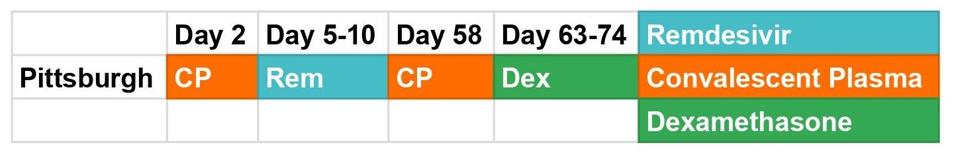
Figure 1. A graphic depiction of the Pittsburgh patient’s treatment course.
“RECURRENT DELETIONS IN THE SARS-COV-2 SPIKE GLYCOPROTEIN DRIVE ANTIBODY ESCAPE” HTTPS://SCIENCE.SCIENCEMAG.ORG/CONTENT/EARLY/2021/02/02/SCIENCE.ABF6950
Like the London and Boston patients, while hospitalized the Pittsburgh patient received a multiplicity of experimental therapies, including the steroid dexamethasone, the antiviral drug remdesivir, and two rounds of convalescent serum, or preparations of highly neutralizing antibodies collected from recovered Covid-19 patients (Figure 1). These treatments put pressure on the virus to evolve—and evolve it did, though in ways that diverge from the cases of other persistently infected patients. What remained consistent across all three, however, is that these mutations evidently gave SARS-CoV-2 some sort of advantage, or it wouldn’t have persisted so long.
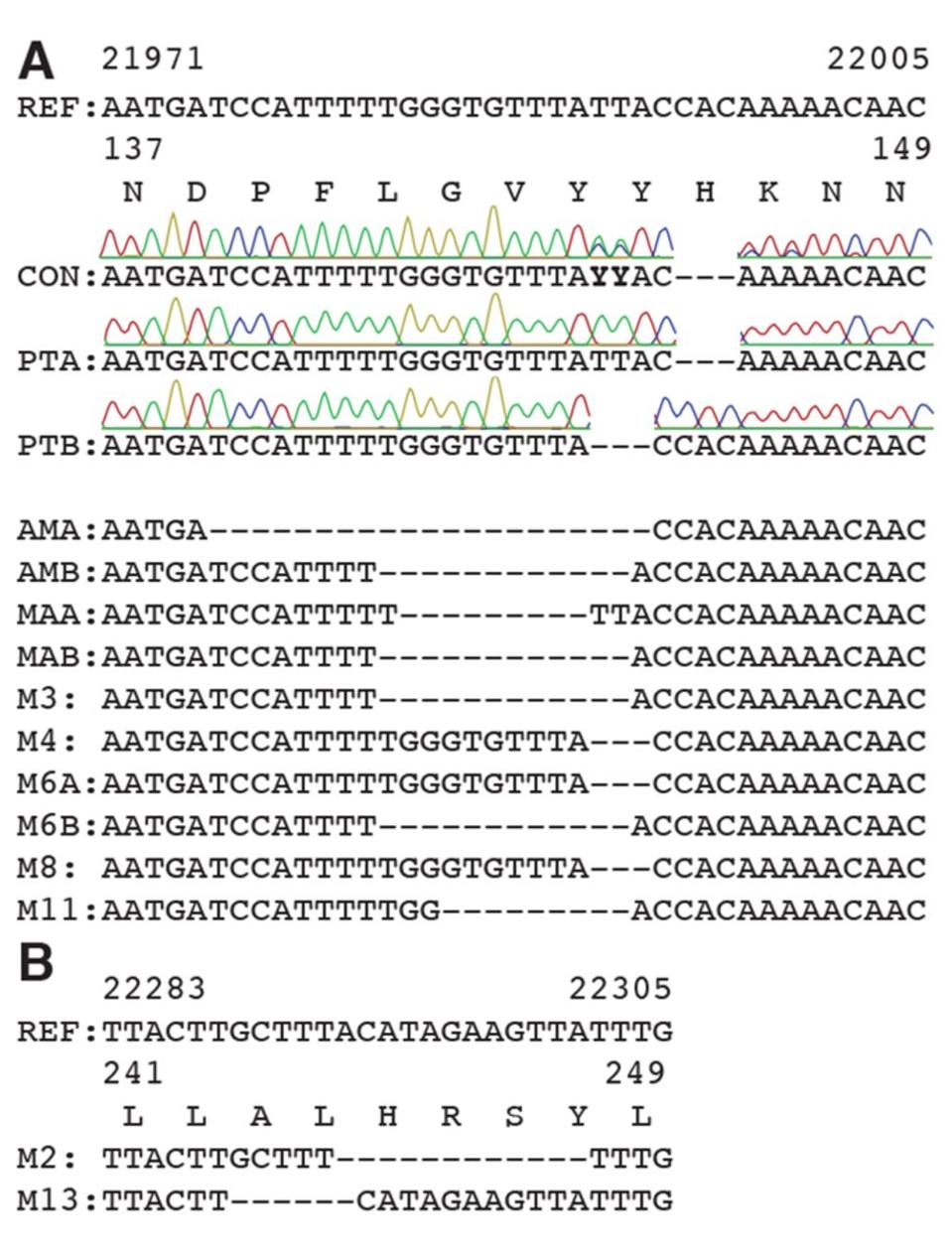
Figure 2. Deletions in SARS-CoV-2 spike arise during persistent infections of immunosuppressed patients.
SOURCE: “RECURRENT DELETIONS IN THE SARS-COV-2 SPIKE GLYCOPROTEIN DRIVE ANTIBODY ESCAPE” HTTPS://SCIENCE.SCIENCEMAG.ORG/CONTENT/EARLY/2021/02/02/SCIENCE.ABF6950
Most of the mutations of note in the London and Boston patients occurred in the spike protein’s receptor binding domain, the target for many drugs and vaccines, as well as the furin cleavage site. Almost all were also point mutations, changes that affected only one amino acid at a time, with the exception of a few deletions. This wasn’t so for the Pittsburgh patient, who had far more deletions in their viral genomes than any other kind of mutation. Additionally, the vast majority were clustered in the N-terminal domain: Δ69-70, Δ141-144, Δ144-145, Δ146, Δ210, and Δ243-244 (Figures 2, 3, 4). If there were mutations in other regions of the virus, they weren’t explicitly stated in the study.
The absence of mutations in the receptor binding domain makes the Pittsburgh patient an outlier. Yet the end result is ultimately the same—the virus persists far longer than normal. Most of the adverse outcomes we’ve attributed to the new SARS-CoV-2 variants—increased transmissibility, immune evasion, and so on—we trace back to point mutations located in the receptor binding domain. But these variants also have mutations in the N-terminal domain that we’d be remiss to neglect.
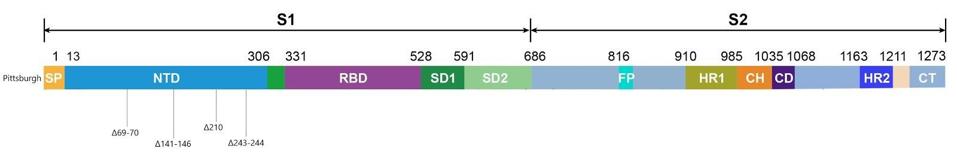
Figure 3. Linear visualization of the Pittsburgh patient’s spike protein genome.
AUTHOR
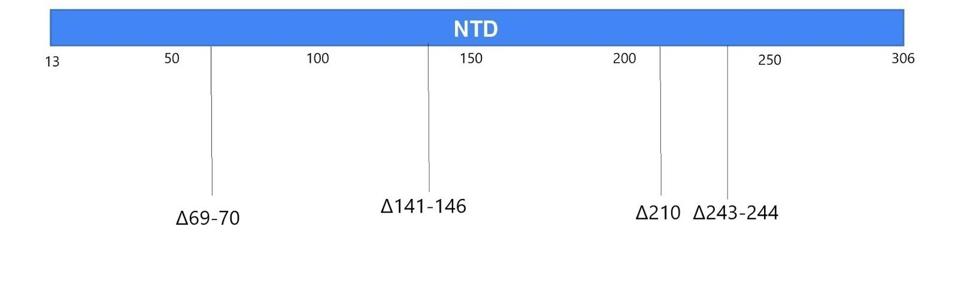
Figure 4. Linear visualization of the N-terminal domain in the spike protein.
AUTHOR
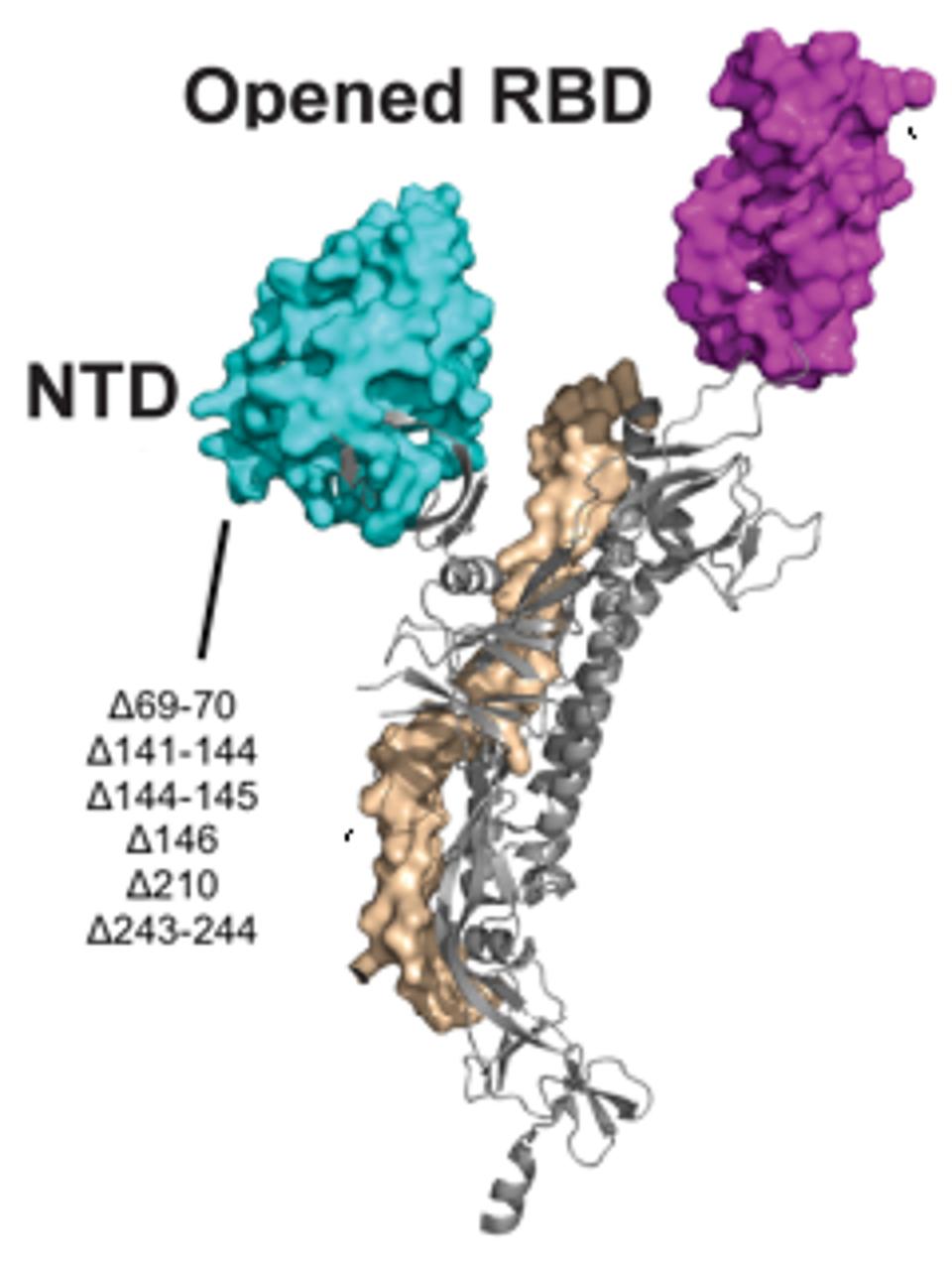
Figure 5. A 3D visualization of the N-terminal domain with notation of the Pittsburgh patient mutations.
“STRUCTURAL AND FUNCTIONAL PROPERTIES OF SARS-COV-2 SPIKE PROTEIN: POTENTIAL ANTIVIRUS DRUG DEVELOPMENT FOR COVID-19” HTTPS://WWW.NATURE.COM/ARTICLES/S41401-020-0485-4 (MODIFIED)
The data on the Pittsburgh patient does indeed suggest that we should pay far more attention to deletions and the N-terminal domain more broadly as potential effectors of change, particularly when it comes to how the virus interacts with the immune system and vice versa. A number of laboratory experiments show that a variety of mutations in the N-terminal domain create resistance to the effects of convalescent sera. The overall function of this globular region of SARS-CoV-2 may be unknown, but I’ve speculated, based on its distinct shape, that in the new variants it could either be increasing affinity as a second receptor or deleting antigenic sites where monoclonal antibodies bind.
Some of the deletions in the Pittsburgh patient we’ve seen before, both at the individual and population level. Δ69-70, for instance, is present in the UK variant, B.1.1.7, while Δ243-244 is a feature of the South Africa variant B.1.153. Not to mention Δ141-146 are nearly the same as deletions found in the Boston patient. When Kevin McCarthy, the lead researcher behind the Pittsburgh patient study, cross-referenced this data with that of GISAID, a genome sequencing database for SARS-CoV-2 and influenza, he found that of all the viruses with deletions in the S protein, 90 percent of those deletions occurred in one of four sites in the N-terminal domain. This means the deletions in McCarthy’s study were prevalent in genomes sequenced all around the world.
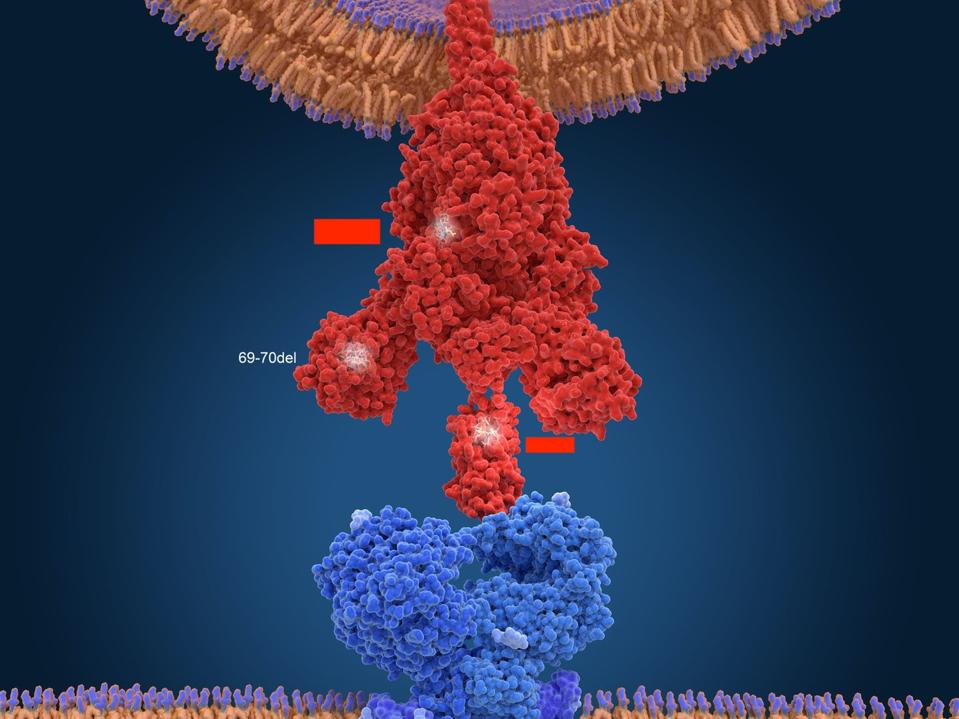
Figure 6. Another 3D visualization of SARS-CoV-2 connecting with an ACE2 receptor.
SCIENCE SOURCE (MODIFIED)
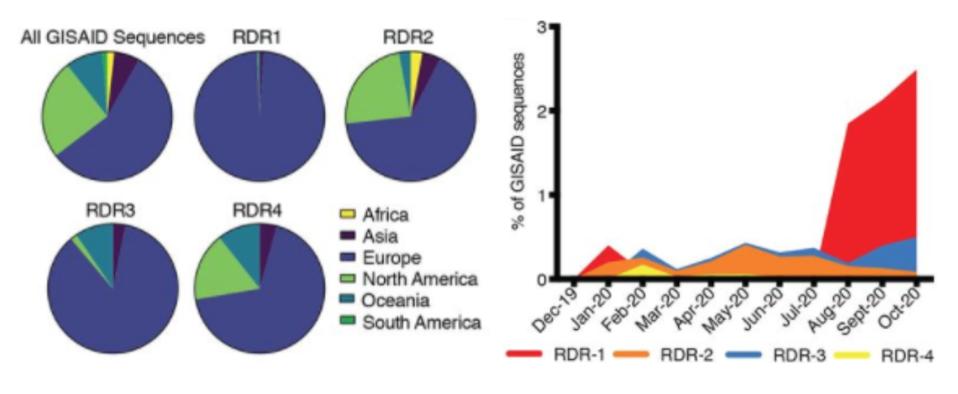
Geographic, genetic, and temporal abundance of RDR variants.
“RECURRENT DELETIONS IN THE SARS-COV-2 SPIKE GLYCOPROTEIN DRIVE ANTIBODY ESCAPE” HTTPS://SCIENCE.SCIENCEMAG.ORG/CONTENT/EARLY/2021/02/02/SCIENCE.ABF6950
The case of the Pittsburgh patient makes one thing clear. If we don’t pay attention to mutations occurring across the entire breadth of the SARS-CoV-2 genome, we risk missing a piece of the puzzle that could prove to be significant for the variants to come. My next piece in this series will be on another persistently infected Covid-19 patient whose mutations might anticipate the ones we see in nature: the Italian patient.
Originally published on Forbes (February 17, 2021)